The analysis and fitting takes place in 384_dilution_analysis.ipynb which should be in a folder for that plate that also includes the excel plate data. Those files reside in a folder entitled "code". Most of the code is in the serology_analysis_functions.py file which relies on NLLSfit.py and FitHill.py for fitting.

Installation of miniconda3 and those packages according to conda installation protocols should be sufficient to run the code. Pandas dataframes with numpy are used for data manipulation and matplotlib and holoviews (via bokeh) are used for visualization purposes. The software is written in python with jupyter notebooks for visualization and report generation. The code should be in a folder entitled "code" at the same level in the folder structure as the folder containing the plate data. No discrimination is made between control and sample rows: such analysis must be done by downstream processing.
Raw absorbance values are assumed to be in an excel worksheet with each well occupying a row. Statistical approaches to analyzing HIV-1 neutralizing antibody assay data. Yu X, Gilbert PB, Hioe CE, Zolla-Pazner S, Self, SG.Monte carlo error analysis is performed by repeating the fit 50 times on data simulated from the best fit and Gaussian random numbers produced according to the distribution of residuals. EC50 values are searched from half of the minimum concentration in the series to 5 times the maximum concentration in the series. The software uses a grid search algorithm with a non-linear search accomplished by multiplying the EC50 value by 1.05 at each iteration. A few notes should be made as to the algorithm used for fitting. Here C max is the maximum concentration in the dilution series. That value is determined as follows:ĪUC = Amp * ( C max - EC50 * ln+ EC50 * ln)
That sensitivity is achieved by measuring the area under the curve (AUC) which is simply the integral over the titration curve (ref?). In particular we are interested in high sensitivity to small amounts of antibody in low titer samples. The EC50 value is in principle a variable related to titer but this value is poorly determined when binding is either saturated (high titer) or doesn't occur at all ( EC50 approaches infinity). Here is a graphical representation of this curve: Here A is the asymptotic amplitude of the signal, EC50 is the relative concentration at which 50% of signal is obtained, c is the relative concentration, and b is the background signal. The methodology behind this software is based on fitting curves to a simple equilibrium binding model: Unfortunately variations in sample-to-sample background levels and the desire to measure low antibody levels make this more complicated. One titrates a serum sample into buffer until signal (absorbance) above background is no longer observed. Serology data analysis is straightforward in principle. Please contact to inquire about non-open source licensing. The license for this software is the GNU GPL version 2 license (see license file) which requires all derivative code to be open source as well. This software is only a tool to aid in that process and its metrics should never be used without visual verification.
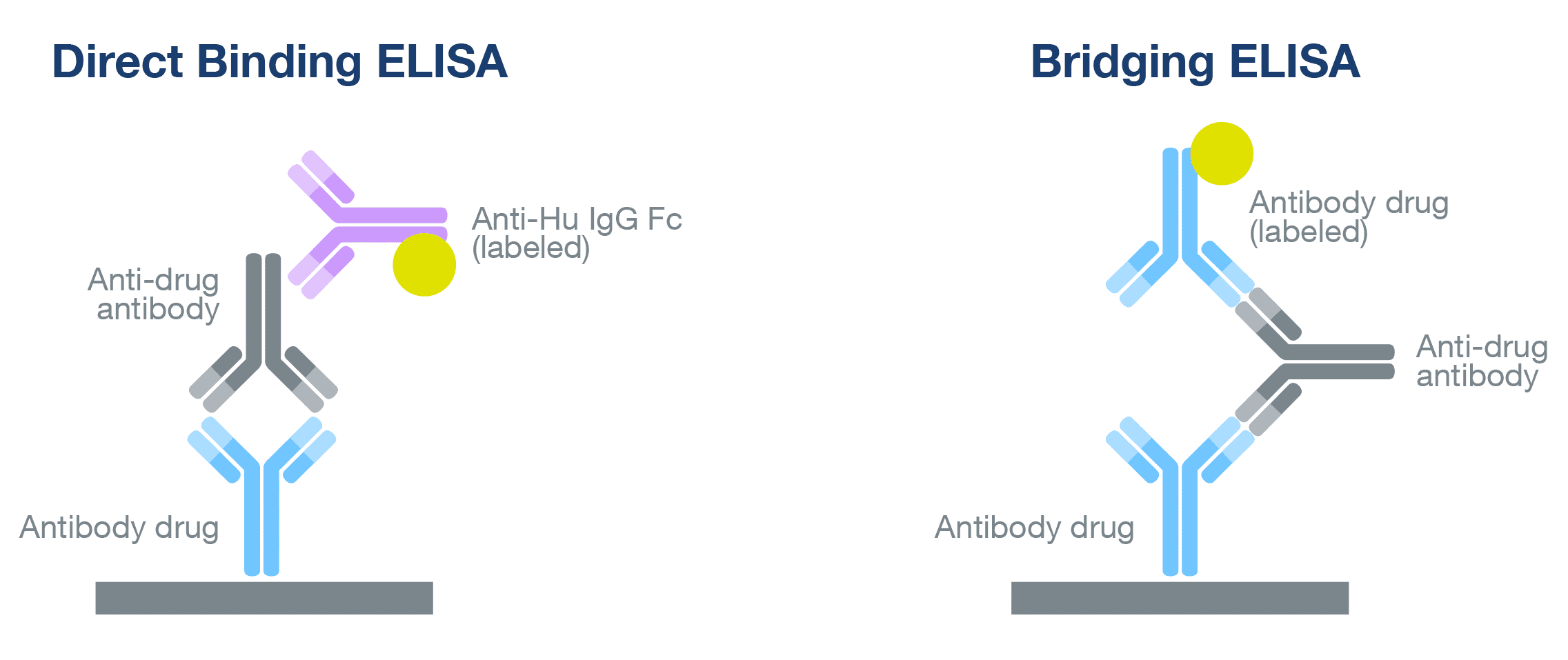

Note that all clinical diagnoses must follow CDC protocols and be verified by visual inspection. This software is provided as is with no guarantee of functionality or support. Serology data analysis with Python for ELISA Antibody Detection
